It has been indicated that traditional sequence comparison methods are linearly sequential in nature and inefficient at identifying CP. In this post-genomic era, the amount of protein structure data is increasing exponentially, and plenty of information should be extractable to reveal the natural prevalence and evolutionary mechanism of CP however, CP search tools are still very rare. In general, previous studies that considered the whole protein as the unit that undergoes CP concluded that CP is rare in nature while those viewing the domain as the unit that undergoes CP suggested CP to be frequent. Besides, because of the disagreement between definitions of CPs, conflicting conclusions can be observed. However, which plays the major role or what proportion each mechanism contributes to the evolution of CPs and protein families remains uncertain. There have been several genetic and evolutionary mechanisms proposed, for instance, duplication/deletion models, duplication-by-permutation models, fusion/fission models, and plasmid-mediated 'cut and paste'. 
CPs can arise from posttranslational modifications but a majority may arise from genetic events. In spite of these interesting properties and applications, there is still much uncertainty about the genetic mechanisms, the evolutionary importance and the natural prevalence of CP. 
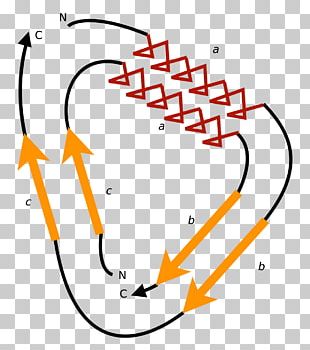
Since CP generally preserves protein structure and function, with sometimes increased stability or activity, it has been applied to trigger crystallization, improve enzyme activities, determine critical elements, and create novel fusion proteins, the tethered sites of which are not confined to the native termini, such as the famous fluorescent calcium sensor. The outcomes have indicated that three-dimensional structure seems remarkably insensitive to CP and CPs generally retain their biological functions, although the structural stabilities, the folding nuclei, transition states or pathways might be altered. To reveal the influences of CP on the structure, function and folding mechanism of proteins, many artificial CPs have been generated, inclusive of trypsin inhibitor, anthranilate isomerase, dihydrofolate reductase, T4 lysozyme, ribonucleases, aspartate transcarbamoylase, the α-spectrin SH3 domain, the Escherichia coli DsbA protein, ribosomal protein S6 and Bacillus β-glucanase. In most of the cases, circular permutants (CPs) have conserved function or enzymatic activity, sometimes with increased functional diversity. Since the first observation of naturally occurring circular permutations in plant lectins, a substantial number of natural examples have been reported, including some bacterial β-glucanases, swaposins, glucosyltransferases, β-glucosidases, SLH domains, transaldolases, C2 domains (for a review, see ), FMN-binding proteins, double-φ β-barrels, glutathione synthetases, DNA and other methyltransferases, ferredoxins, and proteinase inhibitors. It can be visualized as if the original termini of the polypeptide were linked and new ones created elsewhere. Circular permutation (CP) in a protein structure is the rearrangement of the amino acid sequence such that the amino- and carboxy-terminal regions are interchanged. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
